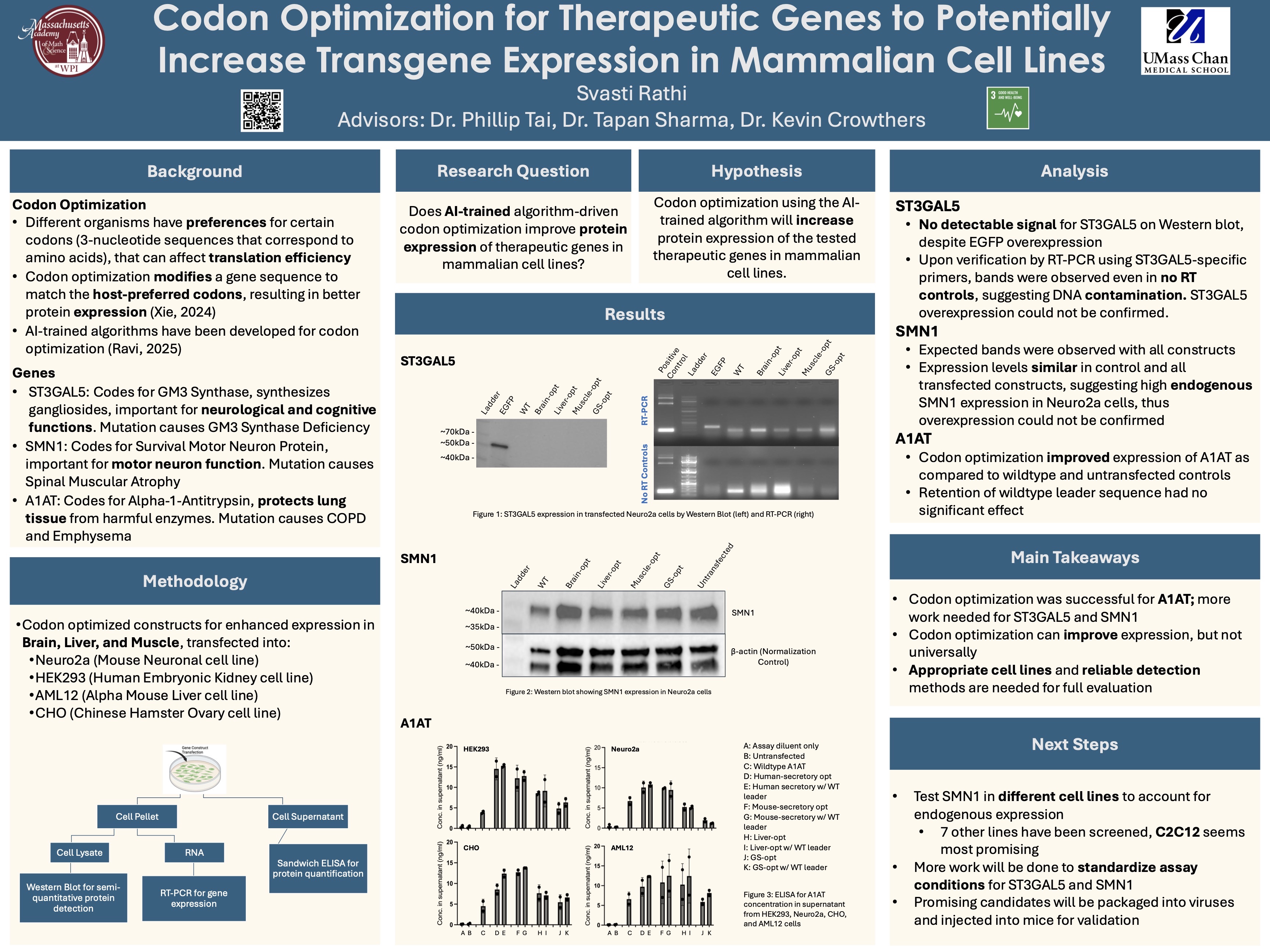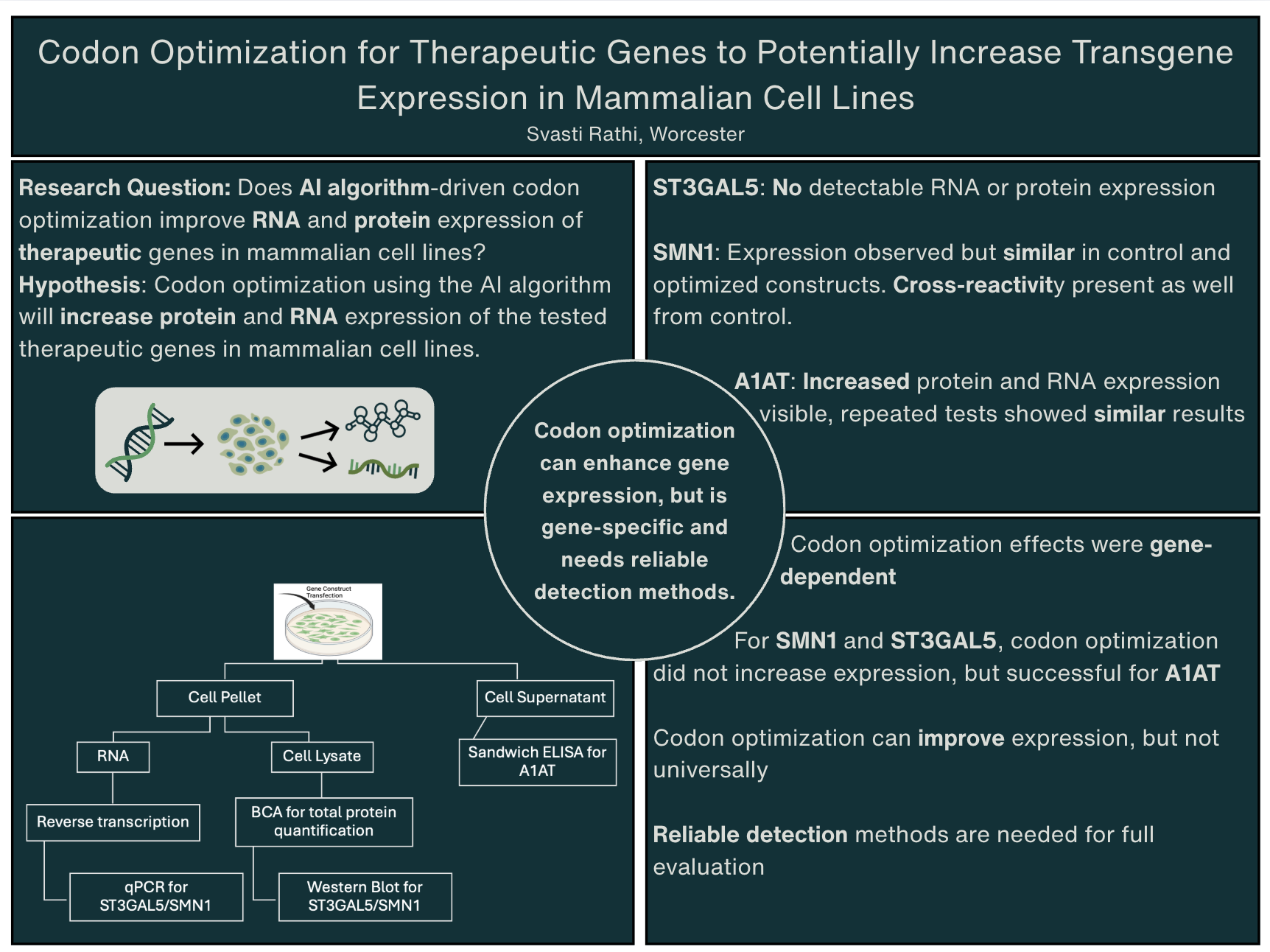

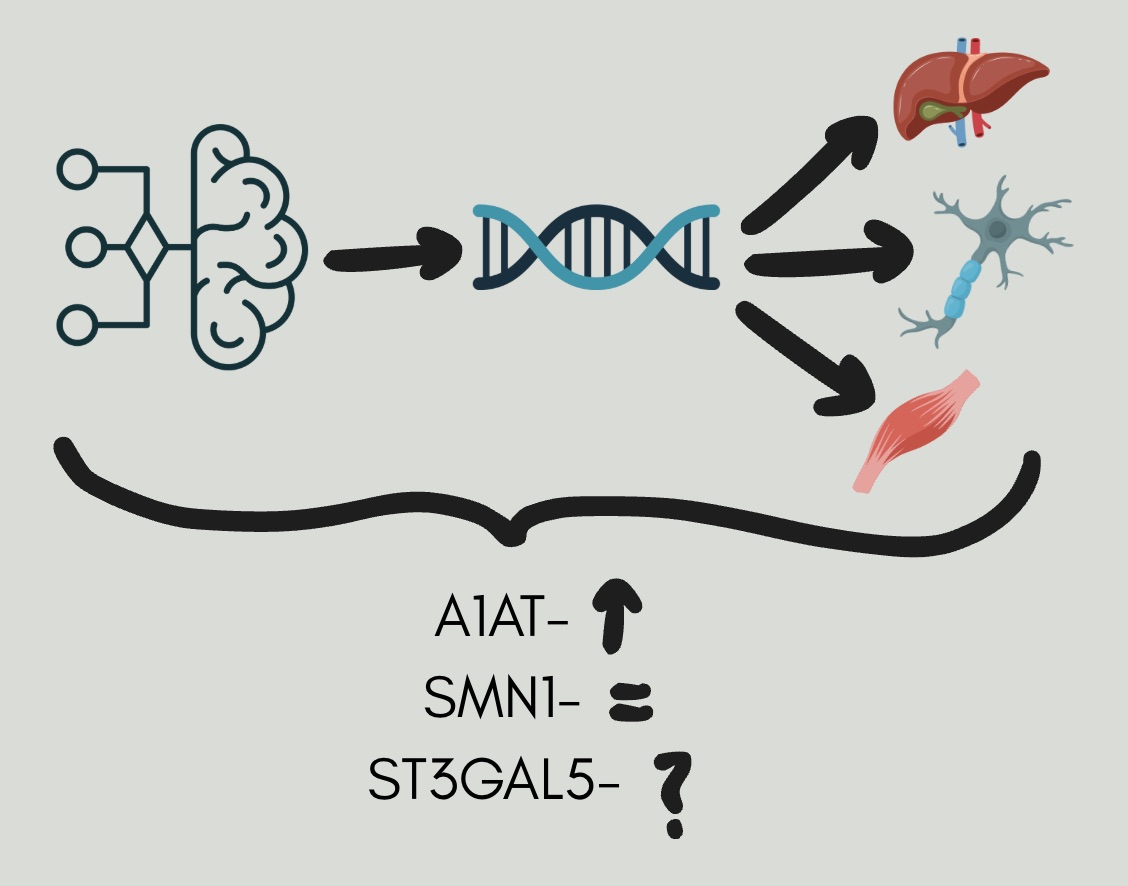
My project investigates whether AI-trained codon optimization can enhance protein expression of therapeutic genes in mammalian cell lines. Codon optimization alters a gene’s DNA sequence to better match host-preferred codons, potentially improving translational efficiency without altering the entire gene sequence or the encoded protein. Three therapeutic genes were studied: ST3GAL5 (important for neurological function), SMN1 (important for motor neuron survival), and A1AT (important for protecting the lungs from harmful enzymes). Codon-optimized constructs targeting brain, liver, and muscle expression were transfected into multiple mammalian cell lines (Neuro2a, HEK293, AML12, CHO). Expression was assessed using Western blotting, RT-PCR, and ELISA, allowing both gene-level and protein-level comparisons between optimized and wild-type constructs.
Codon optimization is a common practice in molecular biology that changes synonymous codon usage to improve translation efficiency without changing the final protein sequence. This project investigates whether codon optimization driven by an AI algorithm developed at the Tai Lab at UMass Chan Medical School can increate RNA and protein expression of therapeutic genes in mammalian cell lines, and whether these results are consistent across various genes. Three therapeutic genes, ST3GAL5, SMN1, and A1AT, were chosen and tested on, using both optimized and non-optimized constructs. After transfections, the RNA and protein levels were assessed using quantitative PCR (qPCR), BCA protein assays, Western blotting, and sandwich ELISA techniques, the latter being used specifically for secreted proteins. Multiple experiments and replications were performed to ensure consistency between results. ST3GAL5 had no detectable RNA or protein expression within the tested conditions. SMN1 expression occurred, but the optimized constructs showed no greater expression compared to the controls. Antibody cross-reactivity also occurred, making the protein analysis more challenging. In contrast, A1AT successfully and consistently showed increased protein expression in the optimized constructs compared to the non-optimized constructs. These findings suggest that codon optimization driven by the AI algorithm can increase transgene expression in mammalian cells but is currently not effective for all genes. The results emphasize the need for gene-specific analysis and tests, along with reliable detection methods for the genes.
Does AI-trained algorithm-driven codon optimization improve protein expression of therapeutic genes in mammalian cell lines?
Codon optimization using the AI-trained algorithm will increase protein expression of the tested therapeutic genes in mammalian cell lines.
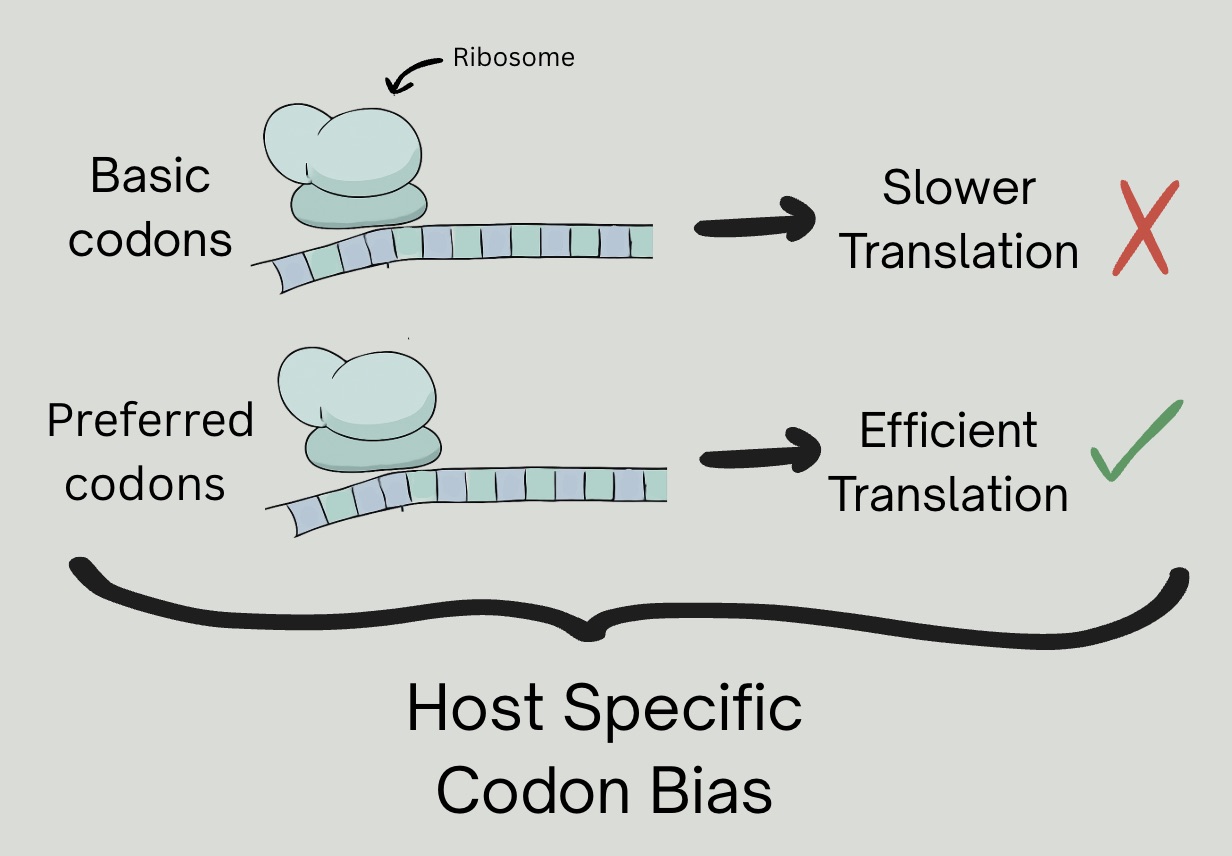
Different organisms have preferences for certain codons (3-nucleotide sequences that correspond to amino acids), that can affect translation efficiency. Codon optimization modifies a gene sequence to match the host-preferred codons, resulting in better protein expression (Xie, 2024). AI-trained algorithms have been developed for codon optimization (Ravi, 2025).
ST3GAL5: Codes for GM3 Synthase, synthesizes gangliosides, important for neurological and cognitive functions. Mutation causes GM3 Synthase Deficiency.
SMN1: Codes for Survival Motor Neuron Protein, important for motor neuron function. Mutation causes Spinal Muscular Atrophy.
A1AT: Codes for Alpha-1-Antitrypsin, protects lung tissue from harmful enzymes. Mutation causes COPD and Emphysema.
The experiments were performed using mammalian cell cultures and general molecular biology techniques. Mouse Neuro2a, human HEK293, Chinese Hamster Ovary (CHO), and Mouse liver (AML12) cells were used to evaluate gene expression in different cellular environments. We designed wild type and optimized constructs for ST3GAL5, SMN1, and A1AT using an AI-based codon optimization algorithm. They were then introduced into cells using transfection reagents.
We assessed the protein expression with Western blot analysis, using antibodies specific for the genes and beta-actin as a normalization control. RNA expression was evaluated using RT-PCR testing (reverse transcriptase PCR) with specific primers for the genes. We quantified secreted A1AT protein levels using Sandwich ELISA assays on the supernatants of the cell cultures. Standard reagents, gel electrophoresis systems, imaging systems and devices, and plate readers were used throughout the study as well.
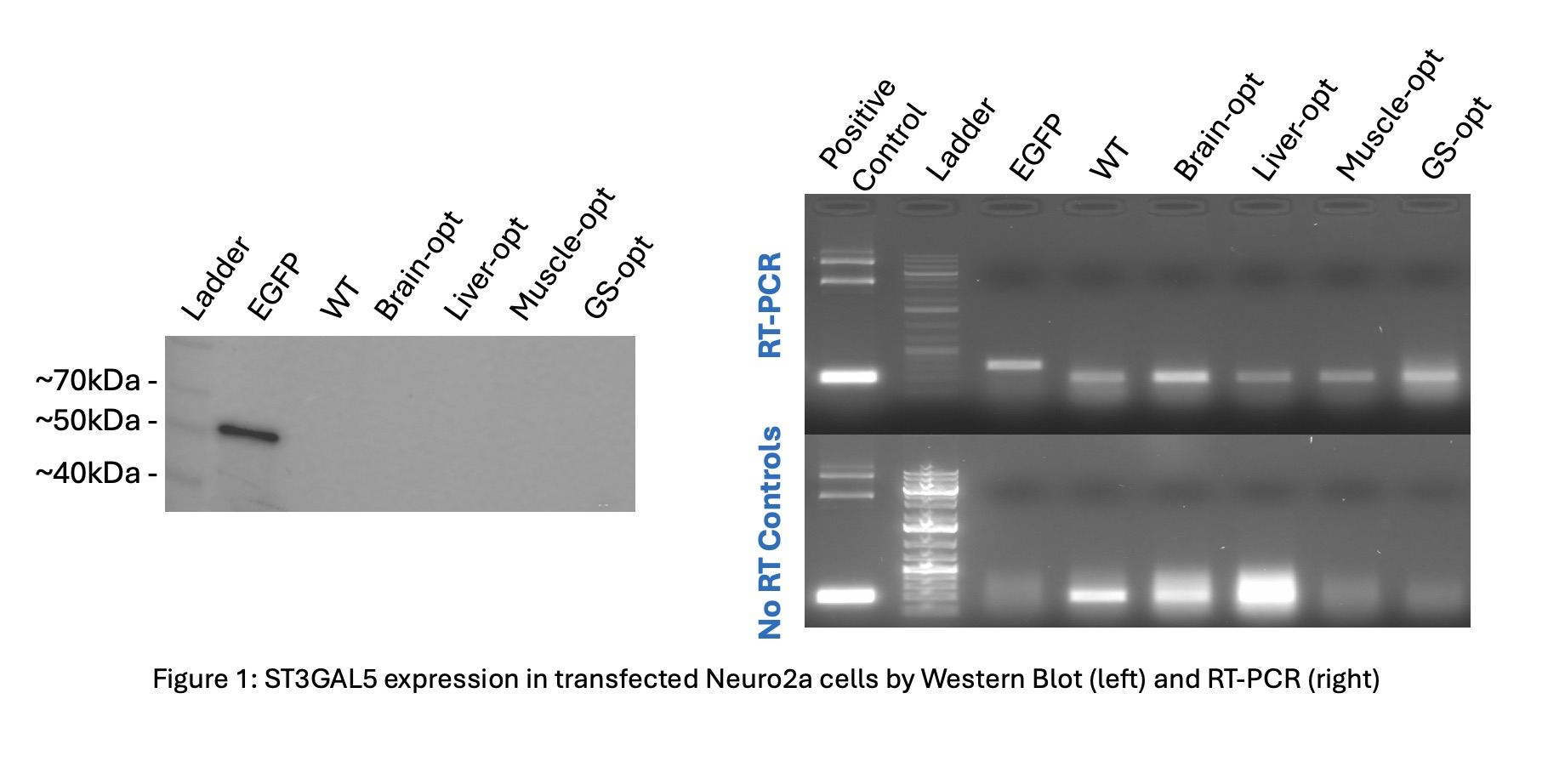
ST3GAL5 Expression: Western blot analysis found no detectable ST3GAL5 protein signal in transfected Neuro2a cells, despite successful transfection controls (Figure 1-left). RT-PCR analysis confirmed amplification in both RT and no-RT samples, indicating DNA amplification instead of true ST3GAL5 overexpression (Figure 1-right).
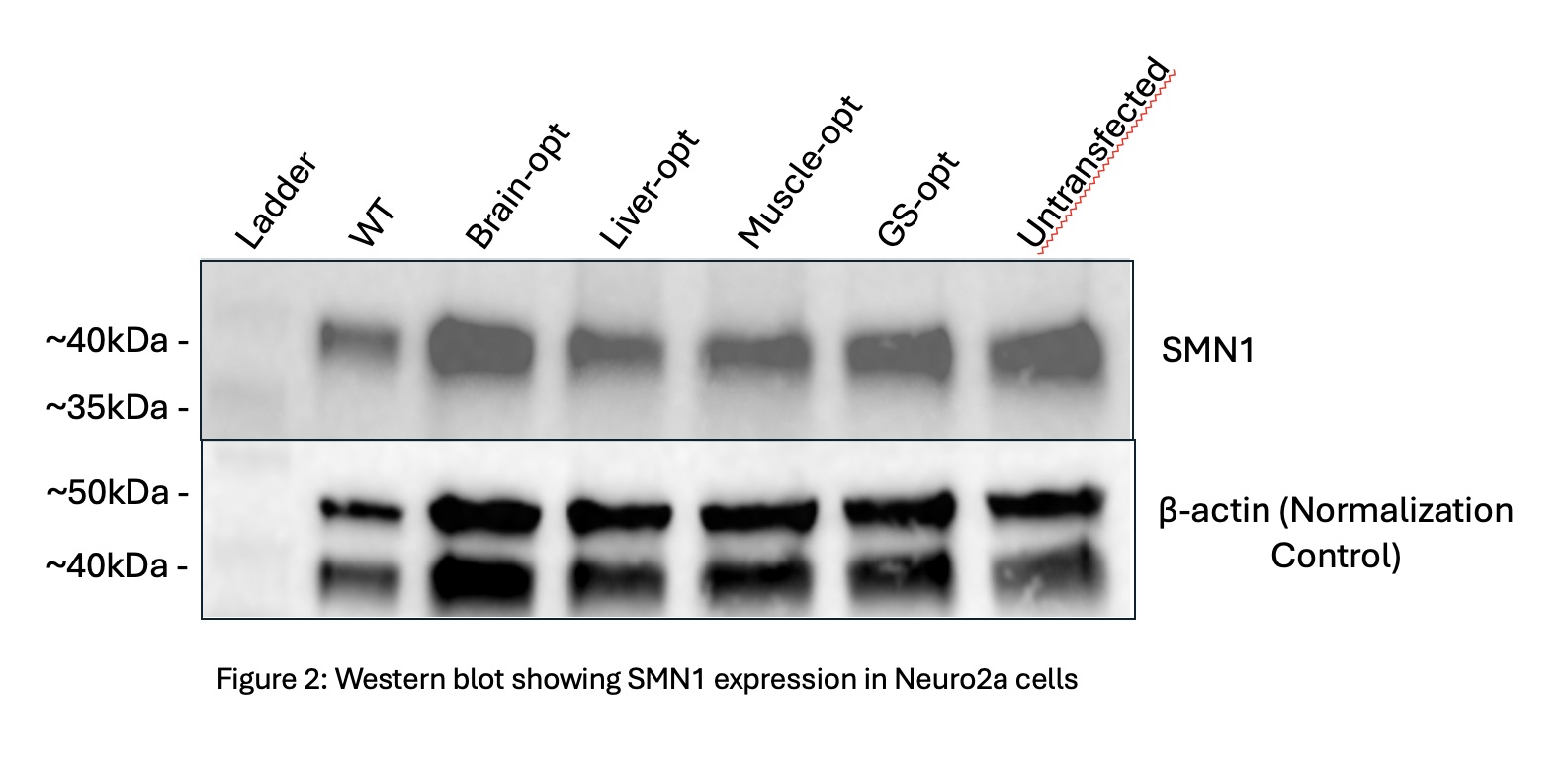
SMN1 Expression: Western blot analysis detected SMN1 protein expression across all constructs, including wild-type and codon-optimized variants (Figure 3). Expression levels looked similar between all constructs, suggesting high basal expression in Neuro2a cells that were hiding the potential overexpression effects on the optimization.
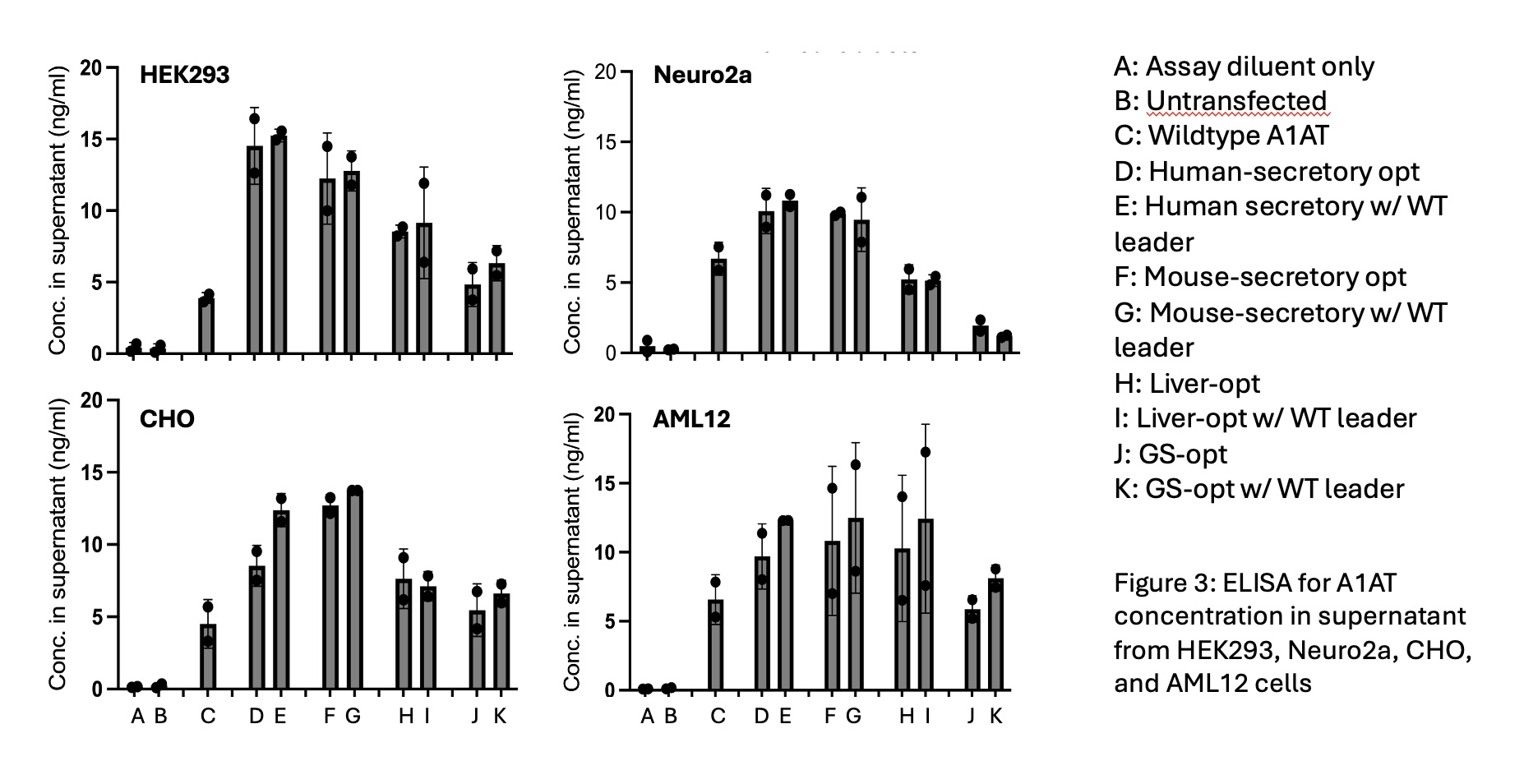
A1AT Expression: ELISA analysis indicated higher A1AT protein concentration in the cell culture supernatants from transfected cells compared to untransfected controls. Codon optimized constructs showed greater A1AT expression than wild-type constructs, indicating sucessful improvement of protein expression due to the optimization algorithm.
This study shows that AI-driven codon optimization can improve transgene expression; however, it is not equally successful for every gene tested so far. Among the genes examined, A1AT had a significant increase in protein expression in the optimized constructs compared to wild-type and untransfected controls, confirming that the optimization techniques were effective for it.
On the other hand, we could not confirm ST3GAL5 overexpression, as no detectable signal appeared in the Western blot. RT-PCR analysis, conducted to verify the Western results, showed amplification in no-RT controls. This suggests DNA contamination, which limited our ability to conclude ST3GAL5 expression. This underscores the need for reliable detection methods and careful validation when assessing outcomes of codon optimization.
For SMN1, expression levels were similar across all constructs and controls. However, we also observed equal expression in the untransfected cells. This likely occurred due to high endogenous SMN1 expression in Neuro2a cells. Consequently, we could not distinguish the effects of the optimization, highlighting the importance of using appropriate cell lines when studying genes with high baseline expression. Overall, these findings support the idea that codon optimization can improve expression, but this has only been seen under specific biological and technical conditions. Effective evaluation will need careful attention to endogenous gene expression, detection sensitivity, and cell line selection. More work is required to fully assess codon optimization strategies for ST3GAL5 and SMN1 using optimized tests and different cell models.