Hi! Welcome to my STEM I page.
STEM I focuses on the Independent Research
Project, like mine, which you can read all about below. We work on
this from the very start of the school year, up until February fair.
Some students move onto WRSEF, or even MSEF/ISEF. While working on
this project, we practice our writing skills with a grant proposal and
a thesis on our project.
A Machine Learning Model for Antibiotic Resistance Gene Forecasting
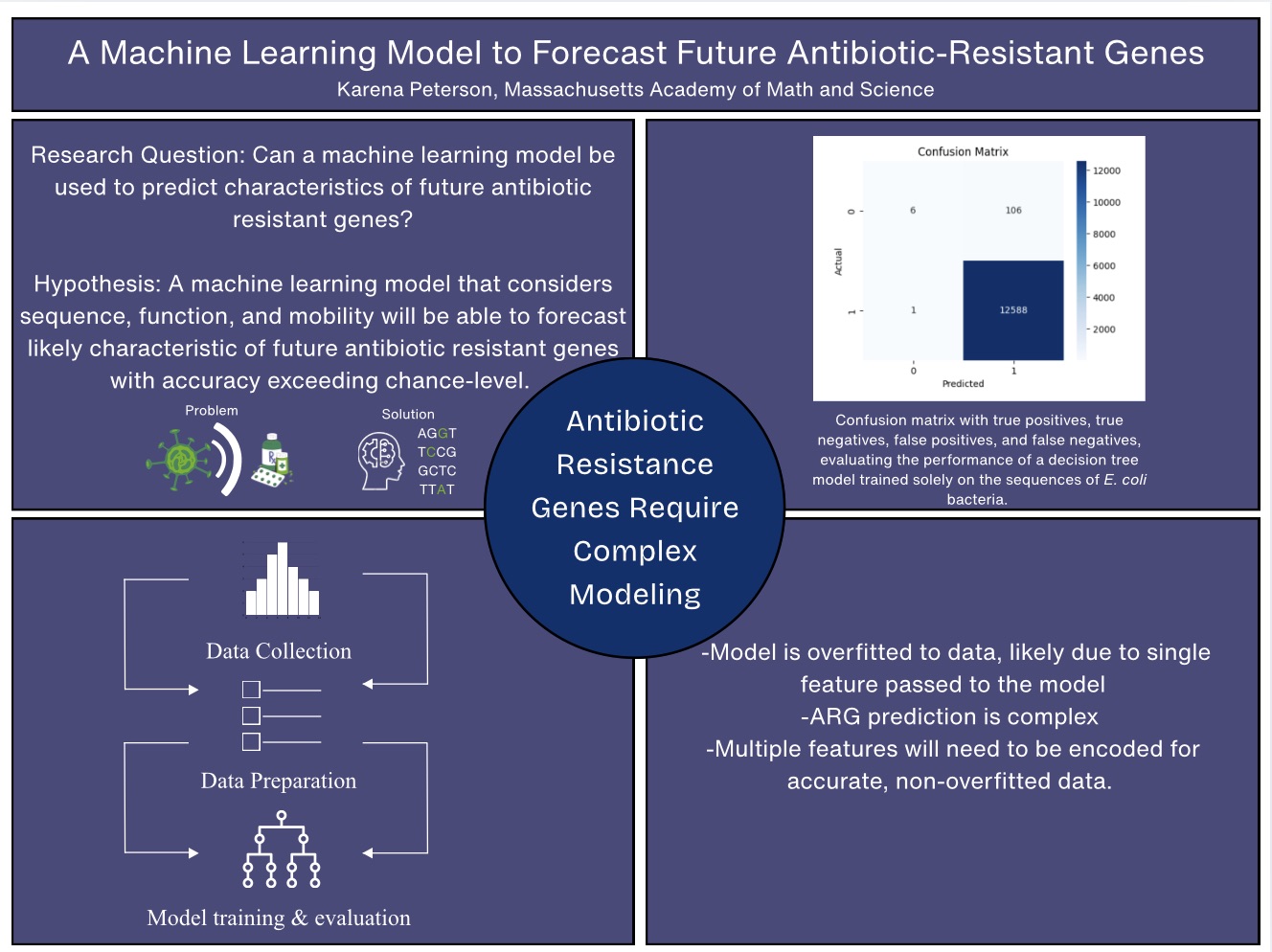
This is a quad chart on my STEM Independent Research Project.
This is the primary focus of the class right now, along with getting
experience pitching and writing about our projects.
A Multi-Layer Machine Learning Model to Forecast Future Antibiotic-Resistant Genes
For my STEM project, I built a multi-layered machine learning model
that evaluates genes before they have mutated for antibiotic resistance.
Research Proposal
Research Question:
Can the patterns of microbial evolution be exploited for a forwards-looking predictive Machine Learning Model?
Hypothesis:
A machine learning model that considers sequence, function, and mobility will be able to forecast likely characteristics of future ARGs with accuracy exceeding chance-level.

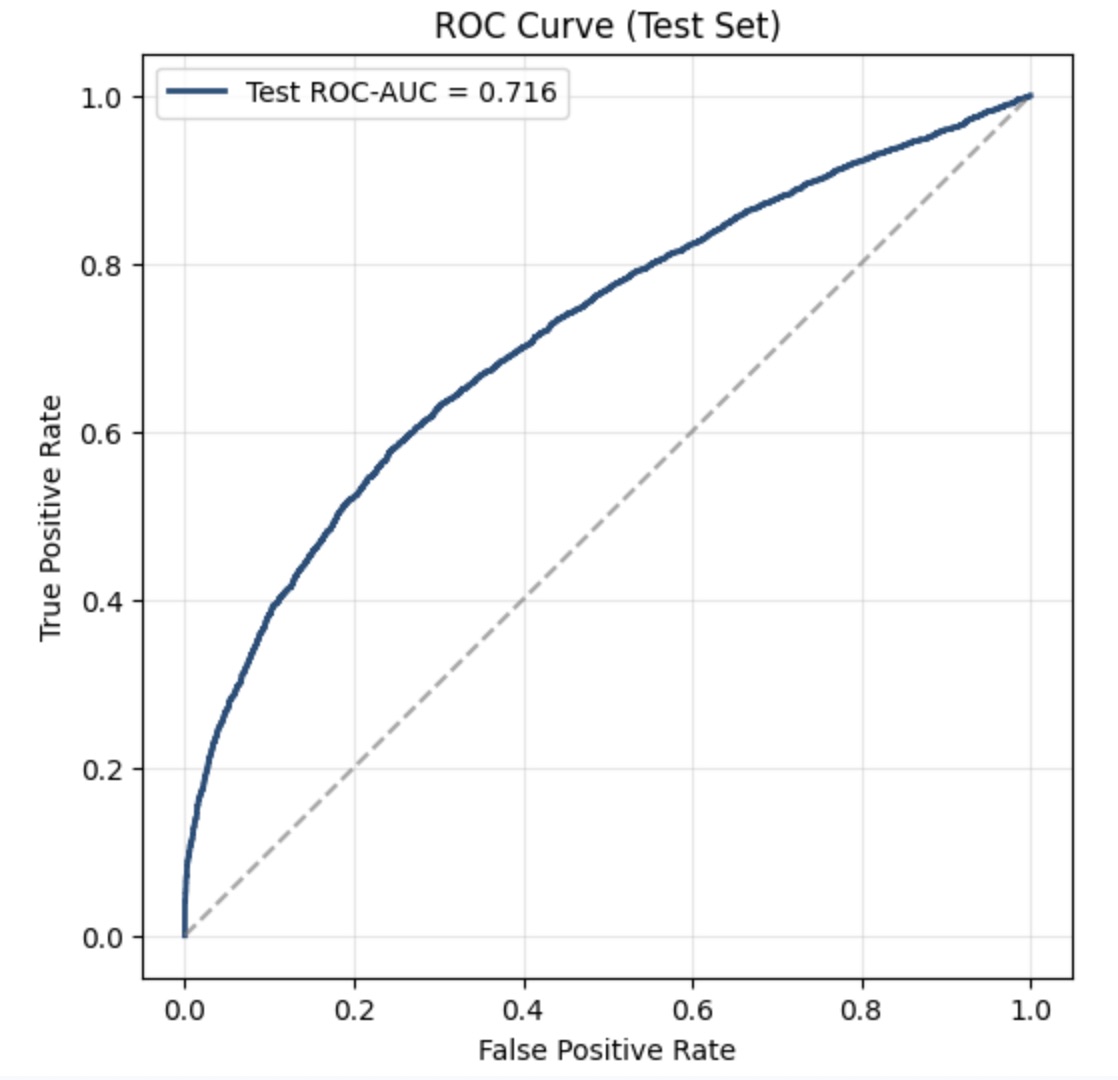
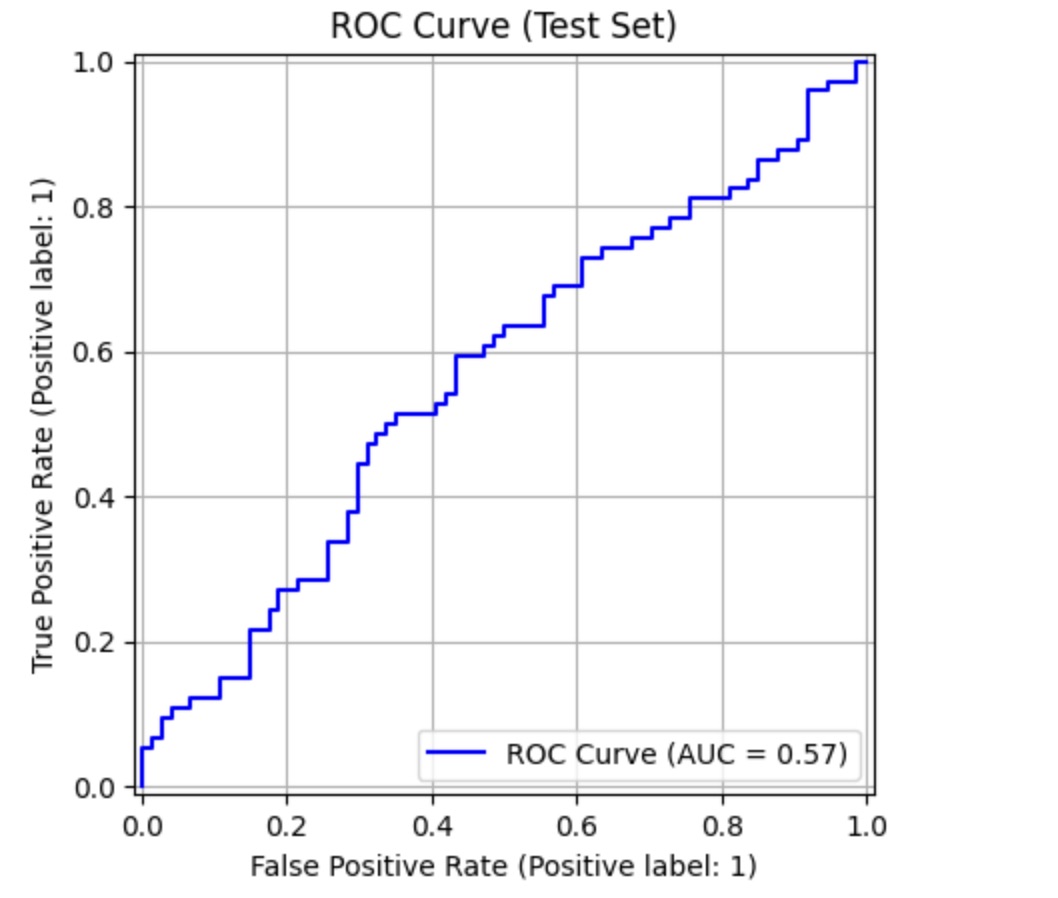
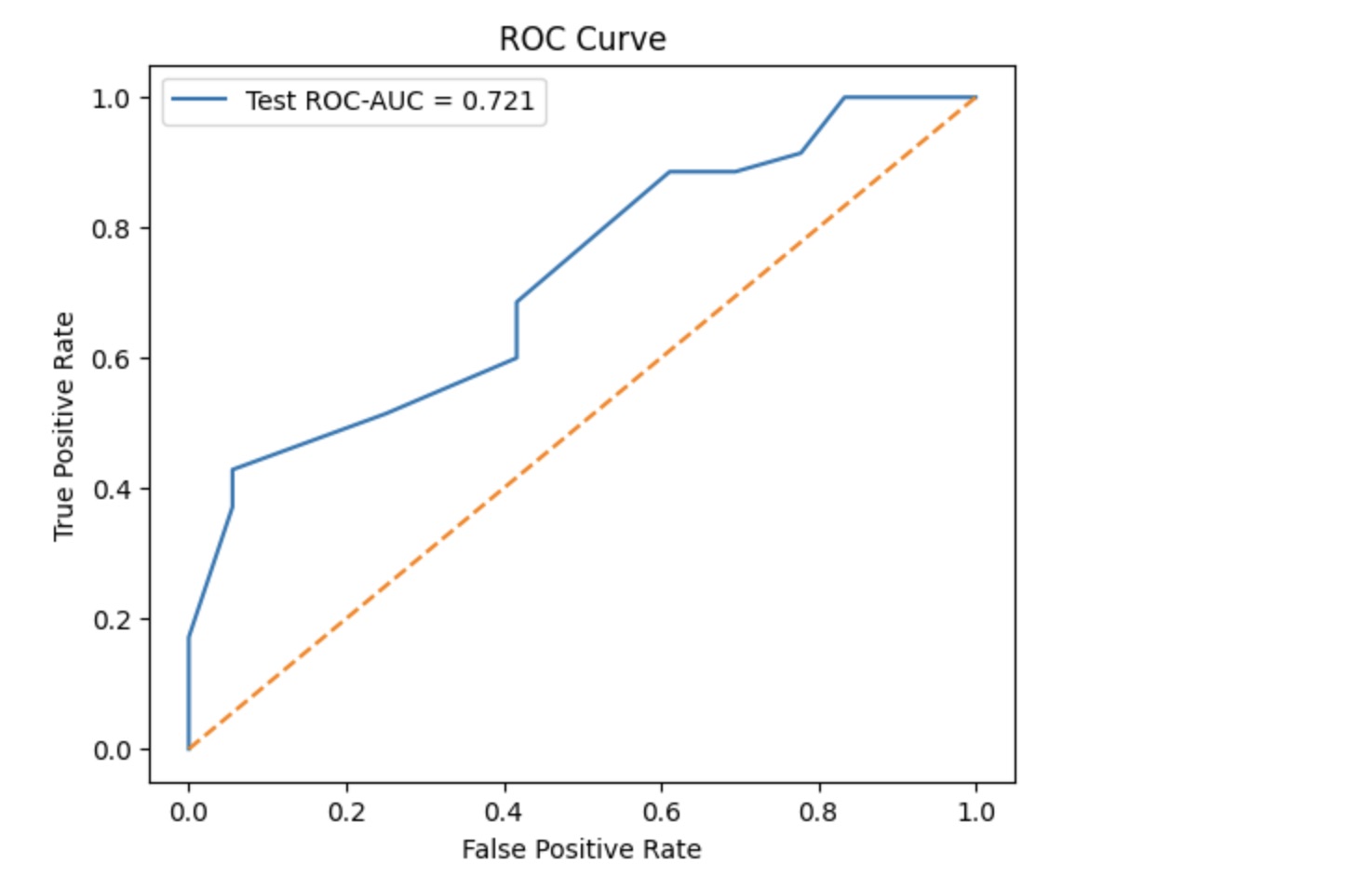
An ROC-AUC demonstrates the ability of the model to distinguish between positives and negatives. The ROC-AUC for the first layer indicates a 71.6% chance that the model will give a higher likelihood of probability to a positive than a negative. For reference, a score of 50% would indicate a model that is randomly guessing between positives and negatives, placing the first layer of the model roughly 20% higher than chance level accuracy. Considering biological restraints and complexity, this is significant. In contrast, the second layer performed much closer to chance level accuracy, with a ROC-AUC score of 57%. This is in contrast to studies that demonstrate a relation between the function of a gene and a mutation in a corresponding gene, suggesting the model may have before badly not due to impossibility but rather small data sets. Finally, the third layer also achieved around 20% above chance-level, with a ROC-AUC score of 72.1%. To further validate these results, a five-fold Cross Validation ROC-AUC was performed. This split the data into five separate training and testing groups, and performed an ROC-AUC test on each of the above. The standard deviation across the five test cases was calculated. Layer one scored 0.0091 SD, layer two scored 0.0304, and layer three scored a 0.0666 SD. The desired SD is a score under 0.05, which indicates low variability and confidence in what the machine learning model learned. While layer one and two each performed well below, layer three demonstrates a slightly above SD.
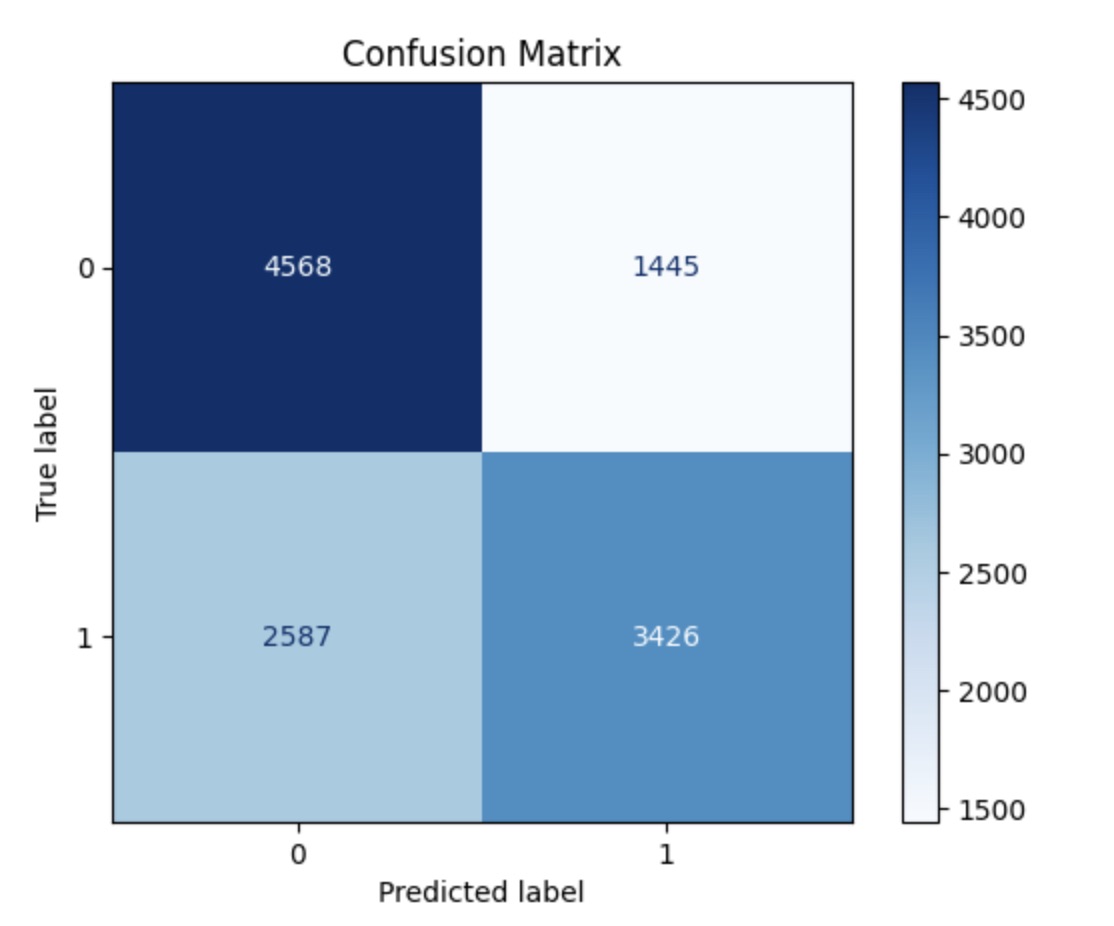
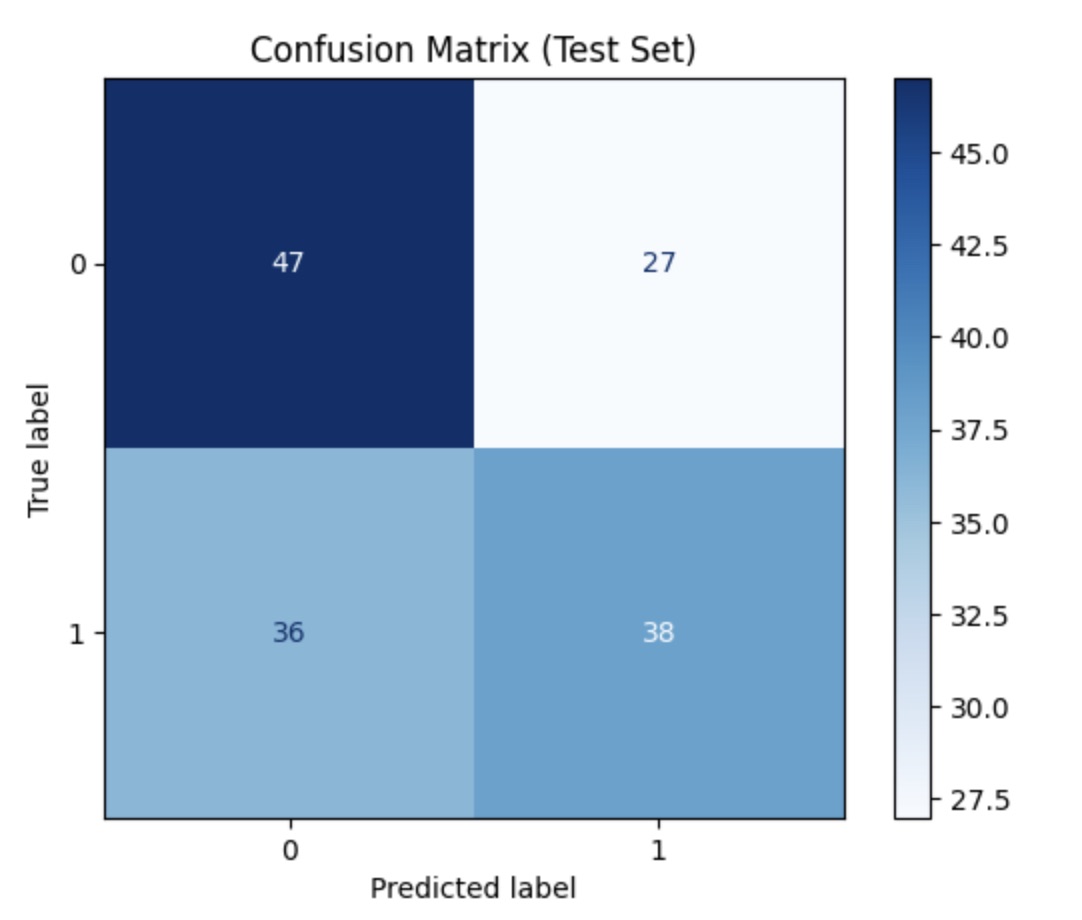
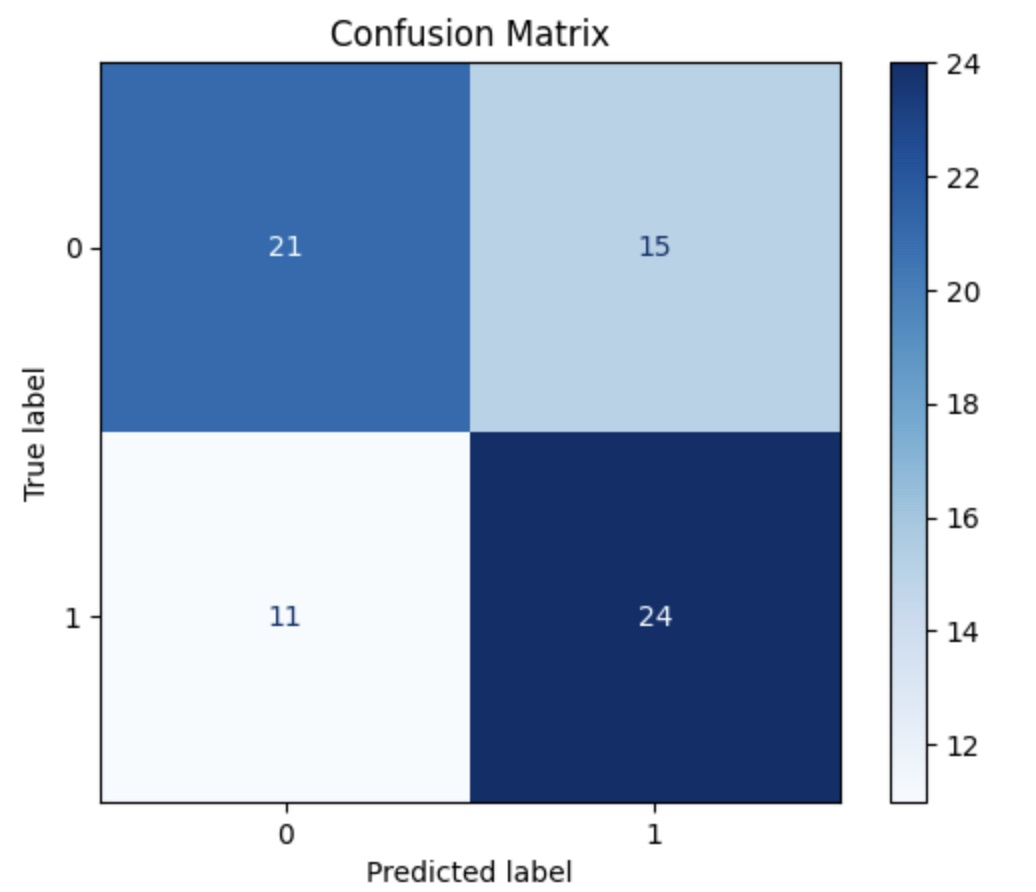
A confusion matrix was also performed for each layer to demonstrate ability of model to classify. The confusion matrix for layer one revealed a strong ability to classify positives correctly, however, with the trade-off of a higher rate of false positives. Similarly, layer two's confusion matrix reveals a strong tendency to classify positive, however, with a high specificity. Finally, layer three's confusion matrix demonstrates a larger tendency to classify as negative than positive.
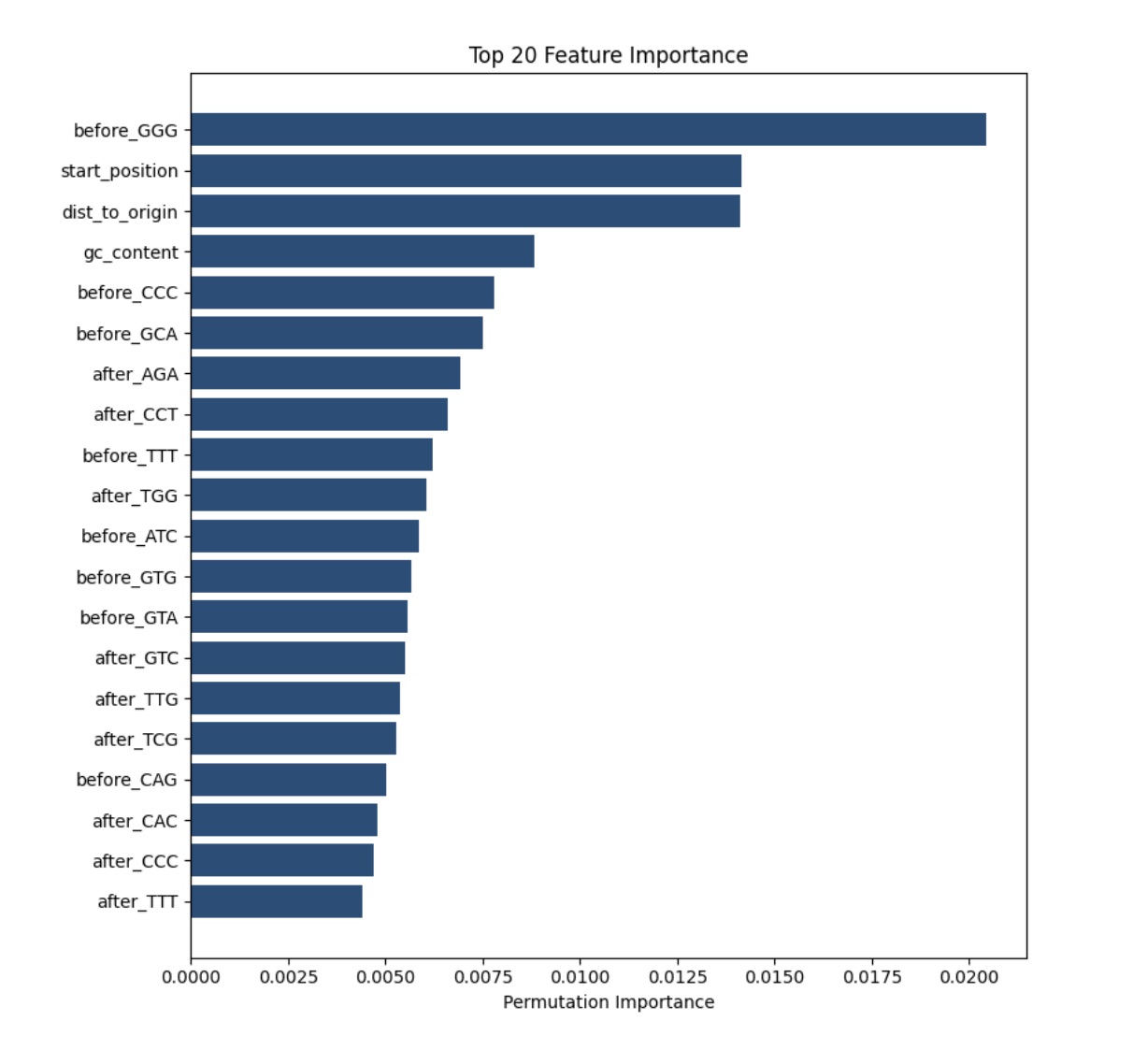
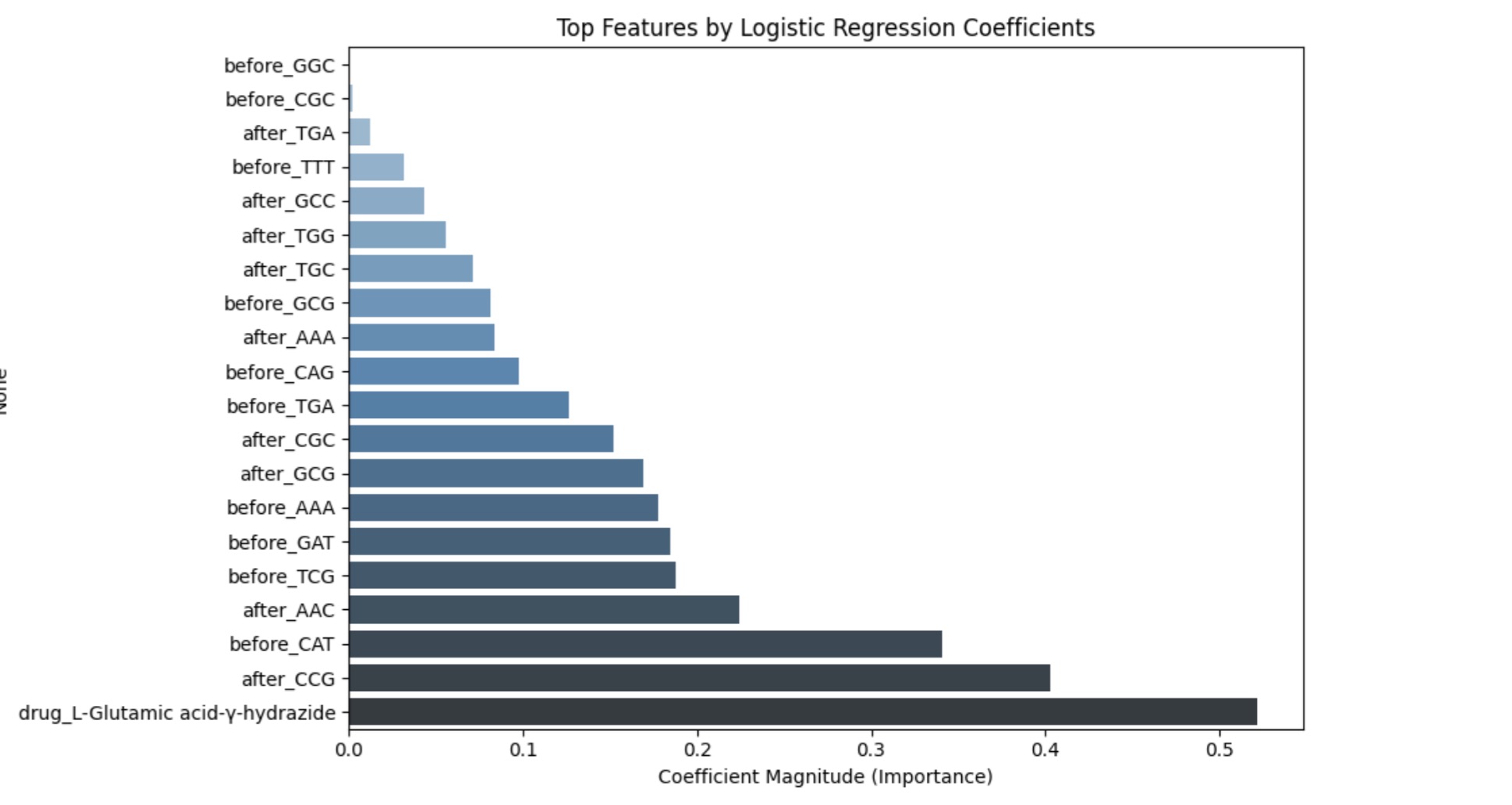
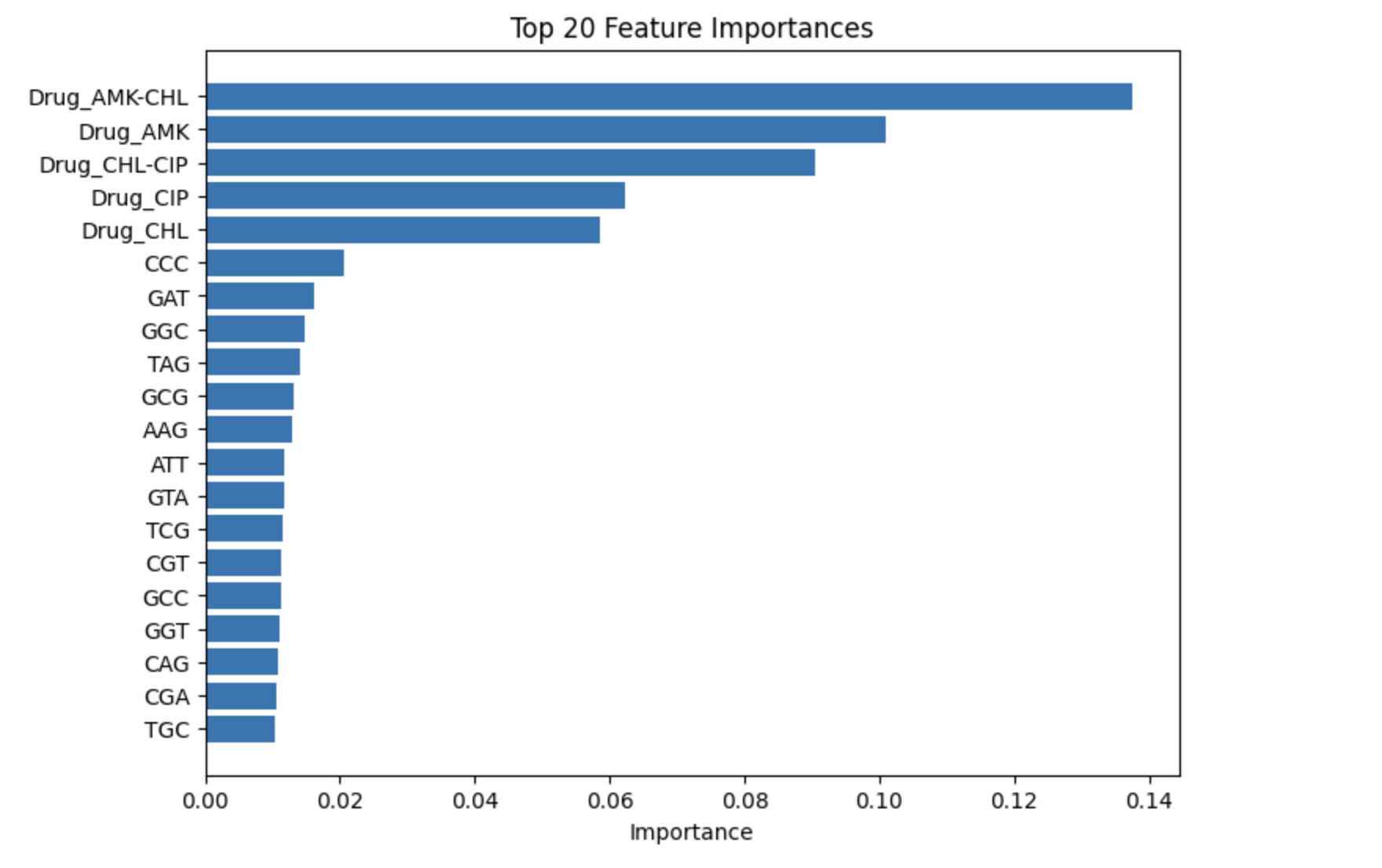
Finally, feature extraction was done for each of the three layers. Importantly, the vast majority of features were k-mer counts, resulting in k-mer counts to consistently appear in the top features. However, it is still important to note which k-mers were deemed more important than the 64 possible k-mer counts. For instance, layer one demonstrates GGG count before the strand in question to be the most important feature. This could indicate GGG to be a signifier for mutation, however, more research is necessary to make a more definitive claim. The next few top features of layer one are not raw sequence information as with k-mers but easily extract-able sequence context. In layer two, a particular drug shows to be the highest feature, which could again indicate that the drug is more likely to lead to AMR encoding mutations. Finally, layer three also ranks all five drugs in the experiment as important factors, demonstrating that certain drugs, depending on their ranking, may be more likely to lead to mutation than others.
Evolution is very complex. However, it appears that evolution can be modeled in increasing specific terms with information gained pre-evolution. Layer one, for instance, was able to achieve an ROC-AUC score that is significantly higher than chance-level accuracy, especially considering biological constraints and complexity. Importantly, easily derivable sequence features other than raw sequences dominate the top features, signaling that the addition of more sequence features may boost performance. In layer two, while much closer to accuracy, a certain method called a Youden's J could find the highest performing split between positives and negatives and may improve the accuracy in layer two. Finally, layer three also showed significant performance considered biological complexity, and may be improved by using a XGBoost tree, essentially an improved Random Forest. Additionally, both layer two and three could benefit from additional data sets, which future work may look towards, as well as the testing of each layer on new data.
Multiple layers of the model show significant potential to be able to predict information about the mutation before it occurs, and it is a promising field that should be further developed. Future work could look at improving upon the accuracy of the model, adding more layers such as mutation function prediction, and the use of a Large Language Model to predict the genome of the mutation.
References:
Alaoui Mdarhri, H., Benmessaoud, R., Yacoubi, H., Seffar, L., Guennouni Assimi, H., Hamam, M., Boussettine, R., Filali-Ansari, N., Lahlou, F. A., Diawara, I., Ennaji, M. M., & Kettani-Halabi, M. (2022). Alternative therapeutic approaches to conventional antibiotics: Advantages, limitations, and potential application in medicine. Antibiotics, 11(12), 1826. https://doi.org/10.3390/antibiotics11121826
Barricklab. (2022). LTEE-Ecoli: Genomics resources for the long-term evolution experiment with Escherichia coli [Source code]. GitHub. https://github.com/barricklab/LTEE-Ecoli
Brown, C. T., Fishwick, L. K., Chokshi, B. M., Cuff, M. A., Jackson, J. M., 4th, Oglesby, T., Rioux, A. T., Rodriguez, E., Stupp, G. S., Trupp, A. H., Woollcombe-Clarke, J. S., Wright, T. N., Zaragoza, W. J., Drew, J. C., Triplett, E. W., & Nicholson, W. L. (2011). Whole-genome sequencing and phenotypic analysis of Bacillus subtilis mutants following evolution under conditions of relaxed selection for sporulation. Applied and Environmental Microbiology, 77(19), 6867–6877. https://doi.org/10.1128/AEM.05272-11
Crozat, E., Philippe, N., Lenski, R. E., Geiselmann, J., & Schneider, D. (2005). Long-term experimental evolution in Escherichia coli. XII. DNA topology as a key target of selection. Genetics, 169(2), 523–532. https://doi.org/10.1534/genetics.104.035717
Despotovic, A., Milosevic, B., Cirkovic, A., Vujovic, A., Cucanic, K., Cucanic, T., & Stevanovic, G. (2021). The impact of COVID-19 on the profile of hospital-acquired infections in adult intensive care units. Antibiotics, 10(10), 1146. https://doi.org/10.3390/antibiotics10101146
Gargate, N., Laws, M., & Rahman, K. M. (2025). Current economic and regulatory challenges in developing antibiotics for Gram-negative bacteria. NPJ Antimicrobials and Resistance, 3(1), 50. https://doi.org/10.1038/s44259-025-00123-1
Good, B. H., & Hallatschek, O. (2018). Effective models and the search for quantitative principles in microbial evolution. Current Opinion in Microbiology, 45, 203–212. https://doi.org/10.1016/j.mib.2018.11.005
Hickman, R. A., Munck, C., & Sommer, M. O. A. (2017). Time‑resolved tracking of mutations reveals diverse allele dynamics during Escherichia coli antimicrobial adaptive evolution to single drugs and drug pairs. Frontiers in Microbiology, 8, Article 893. https://doi.org/10.3389/fmicb.2017.00893
Jezequel, N., Lagomarsino, M. C., Heslot, F., & Thomen, P. (2013). Long-term diversity and genome adaptation of Acinetobacter baylyi in a minimal-medium chemostat. Genome Biology and Evolution, 5(1), 87–97. https://doi.org/10.1093/gbe/evs120
Kallonen, T., Brodrick, H. J., Harris, S. R., Corander, J., Brown, N. M., Martin, V., Peacock, S. J., & Parkhill, J. (2017). Systematic longitudinal survey of invasive Escherichia coli in England demonstrates a stable population structure only transiently disturbed by the emergence of ST131. Genome Research, 27(8), 1437–1449. https://doi.org/10.1101/gr.216606.116
Knöppel, A., Knopp, M., Albrecht, L. M., Lundin, E., Lustig, U., Näsvall, J., & Andersson, D. I. (2018). Genetic adaptation to growth under laboratory conditions in Escherichia coli and Salmonella enterica. Frontiers in Microbiology, 9, 756. https://doi.org/10.3389/fmicb.2018.00756
Lee, C. R., Cho, I. H., Jeong, B. C., & Lee, S. H. (2013). Strategies to minimize antibiotic resistance. International Journal of Environmental Research and Public Health, 10(9), 4274–4305. https://doi.org/10.3390/ijerph10094274
Maeda, T., Iwasawa, J., Kotani, H., Sakata, N., Kawada, M., Horinouchi, T., Sakai, A., Tanabe, K., & Furusawa, C. (2020). High-throughput laboratory evolution reveals evolutionary constraints in Escherichia coli. Nature Communications, 11(1), 5970. https://doi.org/10.1038/s41467-020-19713-w
Naddaf, M. (2024). 40 million deaths by 2050: Toll of drug-resistant infections to rise by 70%. Nature, 633, 747–748. https://doi.org/10.1038/d41586-024-03033-w
National Center for Biotechnology Information. (1998). Escherichia coli str. K‑12 substr. MG1655, complete genome (GenBank accession U00096.3) [Genome sequence]. NCBI. https://www.ncbi.nlm.nih.gov/nuccore/U00096
National Center for Biotechnology Information. (2009). Escherichia coli B str. REL606, complete genome (RefSeq accession NC_012967.1) [Genome sequence]. NCBI. https://www.ncbi.nlm.nih.gov/nuccore/NC_012967.1
National Center for Biotechnology Information. (2011). Escherichia coli str. K‑12 substr. MDS42, complete genome (RefSeq accession AP012306.1) [Genome sequence]. NCBI. https://www.ncbi.nlm.nih.gov/nuccore/AP012306.1
Patra, M., Gupta, A. K., Kumar, D., & Kumar, B. (2025). Antimicrobial resistance: A rising global threat to public health. Infection and Drug Resistance, 18, 5419–5437. https://doi.org/10.2147/IDR.S530557
Rannon, E., Shaashua, S., & Burstein, D. (2025). DRAMMA: A multifaceted machine learning approach for novel antimicrobial resistance gene detection in metagenomic data. Microbiome, 13(1). https://doi.org/10.1186/s40168-025-02055-4
Robillard, D. W., Sundermann, A. J., Raux, B. R., & Prinzi, A. M. (2024). Navigating the network: A narrative overview of AMR surveillance and data flow in the United States. Antimicrobial Stewardship & Healthcare Epidemiology, 4(1), e55. https://doi.org/10.1017/ash.2024.64
Salam, M. A., Al-Amin, M. Y., Salam, M. T., Pawar, J. S., Akhter, N., Rabaan, A. A., & Alqumber, M. A. A. (2023). Antimicrobial resistance: A growing serious threat for global public health. Healthcare, 11(13), 1946. https://doi.org/10.3390/healthcare11131946
Sulayyim, H. J. A., Ismail, R., Hamid, A. A., & Ghafar, N. A. (2022). Antibiotic resistance during COVID-19: A systematic review. International Journal of Environmental Research and Public Health, 19(19), 11931. https://doi.org/10.3390/ijerph191911931
Tang, R., Luo, R., Tang, S., Song, H., & Chen, X. (2022). Machine learning in predicting antimicrobial resistance: A systematic review and meta-analysis. International Journal of Antimicrobial Agents, 60(6), 106684. https://doi.org/10.1016/j.ijantimicag.2022.106684
Ventola, C. L. (2015). The antibiotic resistance crisis: Part 1: Causes and threats. Pharmacy and Therapeutics, 40(4), 277–283. https://pmc.ncbi.nlm.nih.gov/articles/PMC4378521/
World Health Organization. (2020). Global antimicrobial resistance surveillance system (GLASS) report: Early implementation 2020 (CC BY-NC-SA 3.0 IGO). World Health Organization. https://apps.who.int/iris/handle/10665/332081
World Health Organization. (2023, November 21). Antimicrobial resistance. World Health Organization. https://www.who.int/news-room/fact-sheets/detail/antimicrobial-resistance